Xpressor
This work initially stemmed from a related but different question we were working on with Gabriel: “How could we rethink the attention mechanism in transformers to make it more efficient or better?”
Unfortunately, we had to reduce the project’s scope for 2 reasons:
- Time was lacking, Laura wanted us to focus on something more applied to biology and on the next project: scPRINT-2.
- We uncovered not just literature but communities of researchers working on this question 😅
However, we learned a lot and got many ideas. What we had is this notion of an attention matrix that could recursively attend to itself and latent tokens / latent attentions. We initially tried it to reduce attention memory and computational cost. We called it criss-cross attention, it actually worked well and is mentioned in the scPRINT-2 paper!
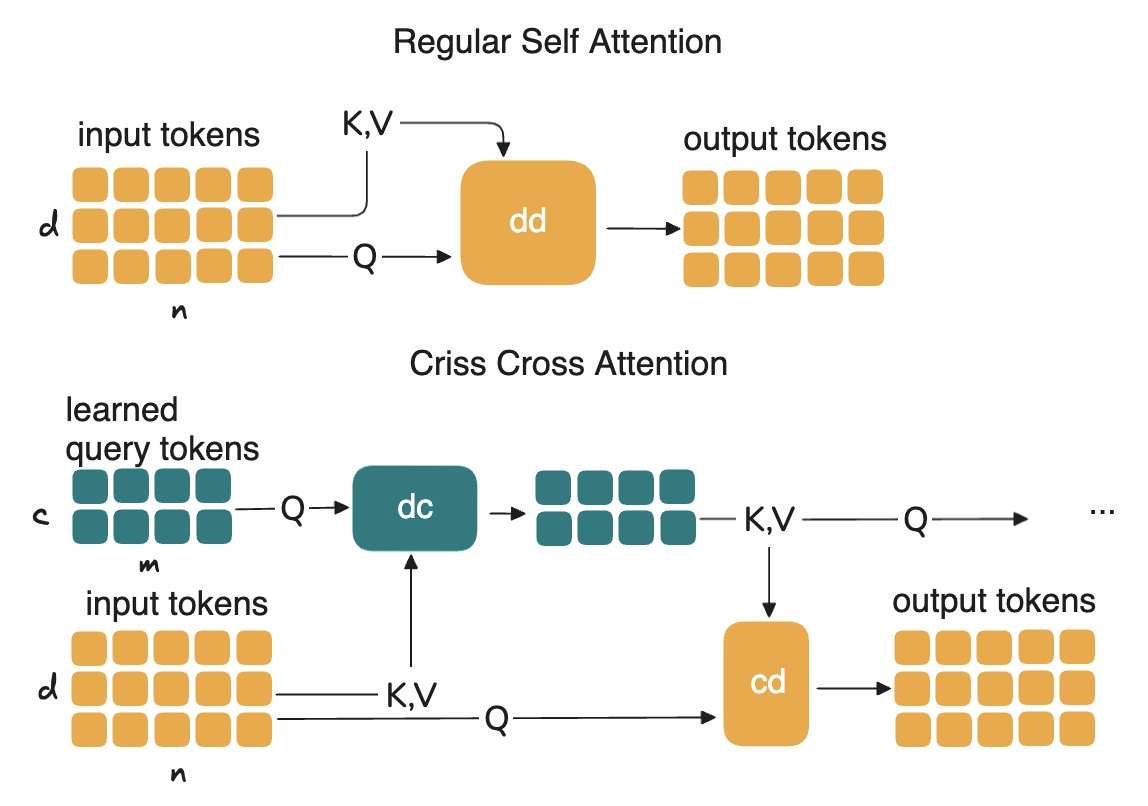
But this had not enough biological applications and Laura prefered to not publish on this in an ML conference. Gabriel felt that we needed more theoretical work to understand the properties of this attention, something of a disagreement between my PIs.
In the end, we merged it with another question we had: “How to make cell embeddings of single-cell foundation models like scPRINT, better. Indeed to make an embedding from a transformer’s input there was 2 main ways: either to use a [CLS] token (class-pooling) or to average all tokens (avg-pooling). Both were not great, and did not allow reconstruction of the model’s input from the embedding, as an auto-encoder would do.
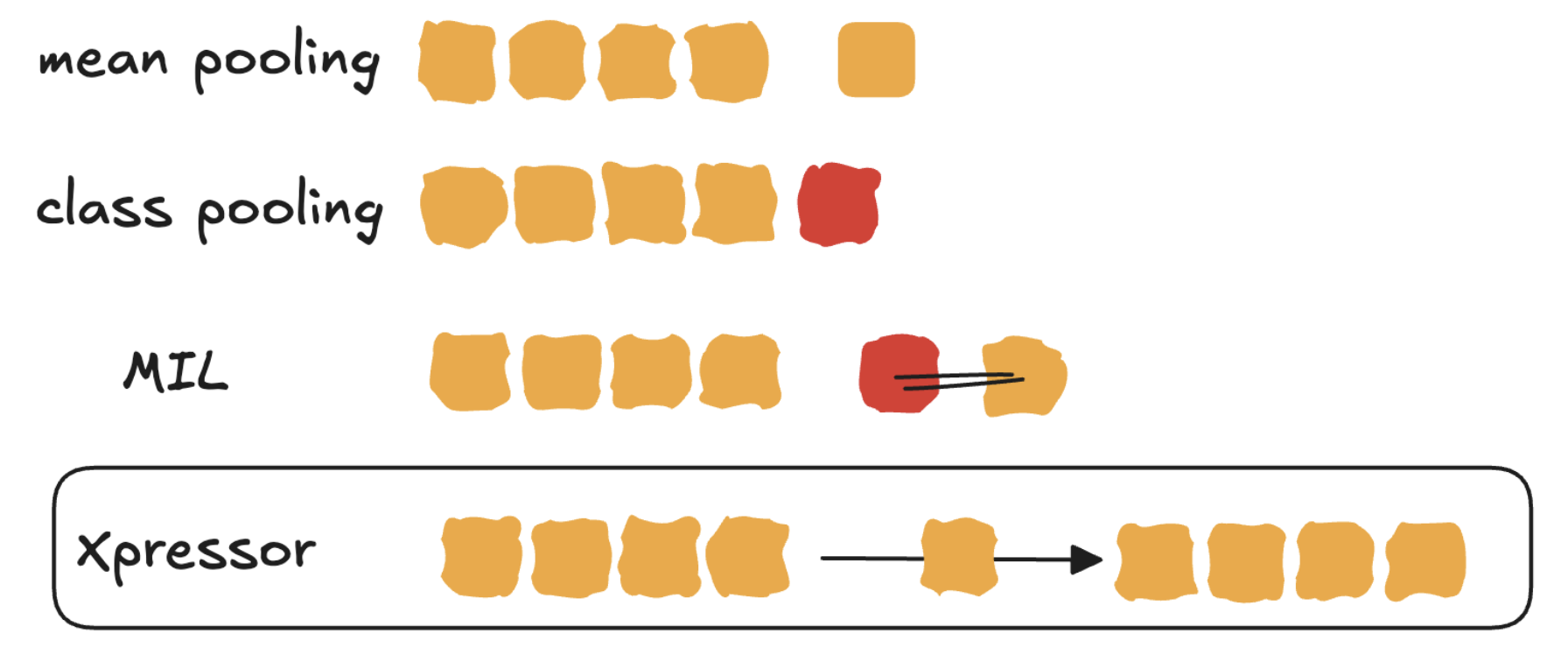
This is how we came up with the idea of Xpressor, a model that would learn to reconstruct its input from a latent embedding, and that would have an attention mechanism that would allow it to attend to itself and to latent tokens. We realized that bringed 2 new possibilities:
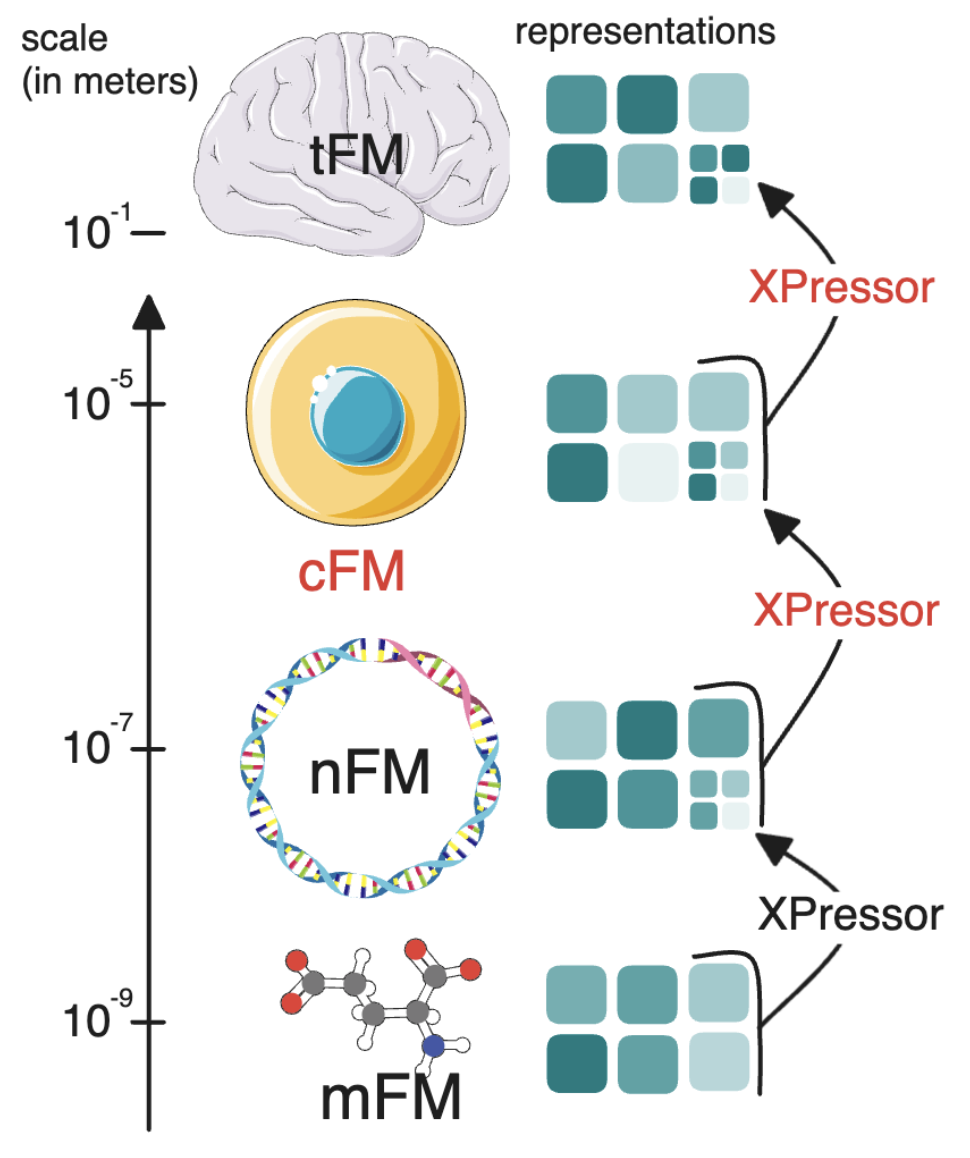
1. Cross scale learning:
extending it outside single cell, it could be a way to learn across biological scales by compressing cells into input tokens for tissues, by compressing genes into input tokens for cells.
2. Parameter efficient fine tuning:
by adding a small transformer (called xpressor blocks), we could be both more expressive but also do a very parameter efficient fine tuning of the xpressor blocks. This was used heavily in scPRINT-2 for batch correction, and cell type prediction, but it can be used to fine tune a model from a scale to another, something we showed in Xpressor by fine-tuning ESM-2 (a protein language model) to generate protein embeddings that were useful for single cell predictions.



Leave a comment