This is an important moment for me, and it seems that many things led me to this point. 😊 I have decided to join Laura Cantini at Institut Pasteur and Gabriel Peyré at ENS ULM for a joint Ph.D. in Computational Biology and Applied Mathematics. 🧬📊 I will try to explain here my decision process for something that can appear as a very sudden change of direction and slightly crazy, given my current position at WhitelabGx.
You can learn more about what I did at WhitelabGx here but I don’t think it would explain much of my decision. I will try to explain here the reasons that led me to this decision and the many lessons I learned along the way. 📚🤔
First, as you might be able to see in this life story of mine, I was always into science and research. I got proposed quite a few Ph.D. positions along the way. At ECE, University of Kent, EPFL, and INSERM. I applied myself to quite a few competitive places. But always found some other directions that I was more inclined to follow. It always seemed like I was pushing it for later or just thinking I could do without it. In a way I did. I went to work at the Broad Institute. Worked on many research projects, published as first-author, and even started working on my own projects. I became a research scientist in companies and even came on to lead scientists themselves during my time at Whitelab.
So research, although not my only passion, has always been very important to me.🧪🔬 It always seemed obvious that I would go on and do a Ph.D. at some point. But why did I wait? Why now? 🤔
The story
I was not looking for any other positions. I had in mind to stay at least a few years at Whitelab Gx, see where it had gone in that time, and then make a decision about my future.
However, I was in contact with Laura Cantini with whom I had discussions about Ph.D. projects before joining Whitelab. At some point, Laura came back to me with this Ph.D. proposal. I spent the good part of a month in this tough dilemma. Thinking about it in all possible ways to try and find a solution that I would not regret in the future. 🤯
I will try to explain below what led me to the decision to take this Ph.D. opportunity.
But still, I did not want to let Whitelab down. during the end of my review period, I asked for it to be extended and explained the reason why. That gave Whitelab 2.5 months to prepare for my departure. Moreover, I presented a proposition to keep an advisory role in the company during my Ph.D. and to spend a week or so preparing my substitute for the position. I also did not take any vacation between my time at Whitelab and the Ph.D. so as to give as much time to Whitelab as I could. 🤝📅
I believe that this goes to show my dedication and interest in the company, its mission, and the respect I have for my colleagues. 🙏🏢
the why
All the opportunities I had were very exciting to me. They were allowing me to learn in some of the best environments. At Broad Institute, I learned how to do research, biology, genomics, and cutting-edge tools and techniques.
At Whitelab I learned how people build companies, how to lead a team, and how to go start from scratch a unit, with a validated plan, product design, and layout a business strategy for it. From working with clients, the BD team and the management.
Why now? Well, there is no perfect time. But I could summarize it into 1 reasons:
- The Ph.D. topic and group which I believe was a great fit for me.
Obviously not everything can be summarized into one reason and not everything was perfect otherwise:
- Whitelab was maybe a bit too early stage for me? At least it is something I thought about sometimes see Whitelab post
- I thought a lot about open source in computational biology and having an impact which I will be able to outline further in this post.
- I also had a couple disillusions during job interviews with some high profile companies a few years back see leaving Broad.
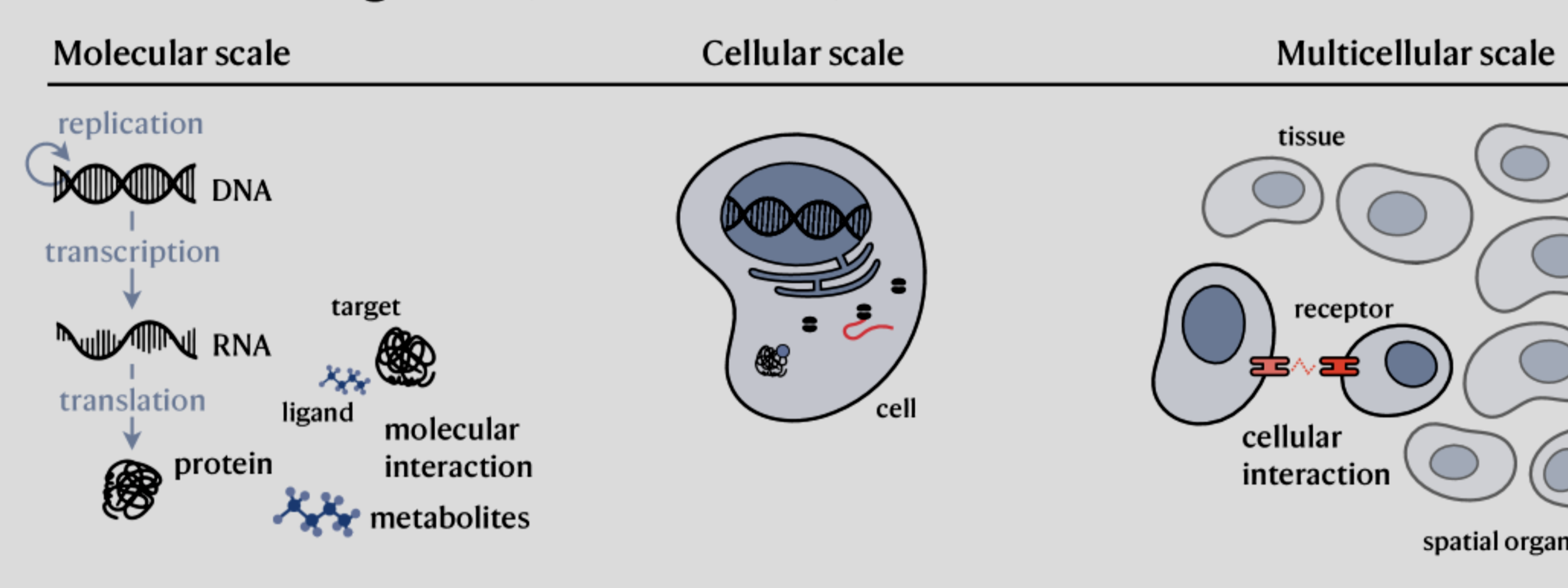
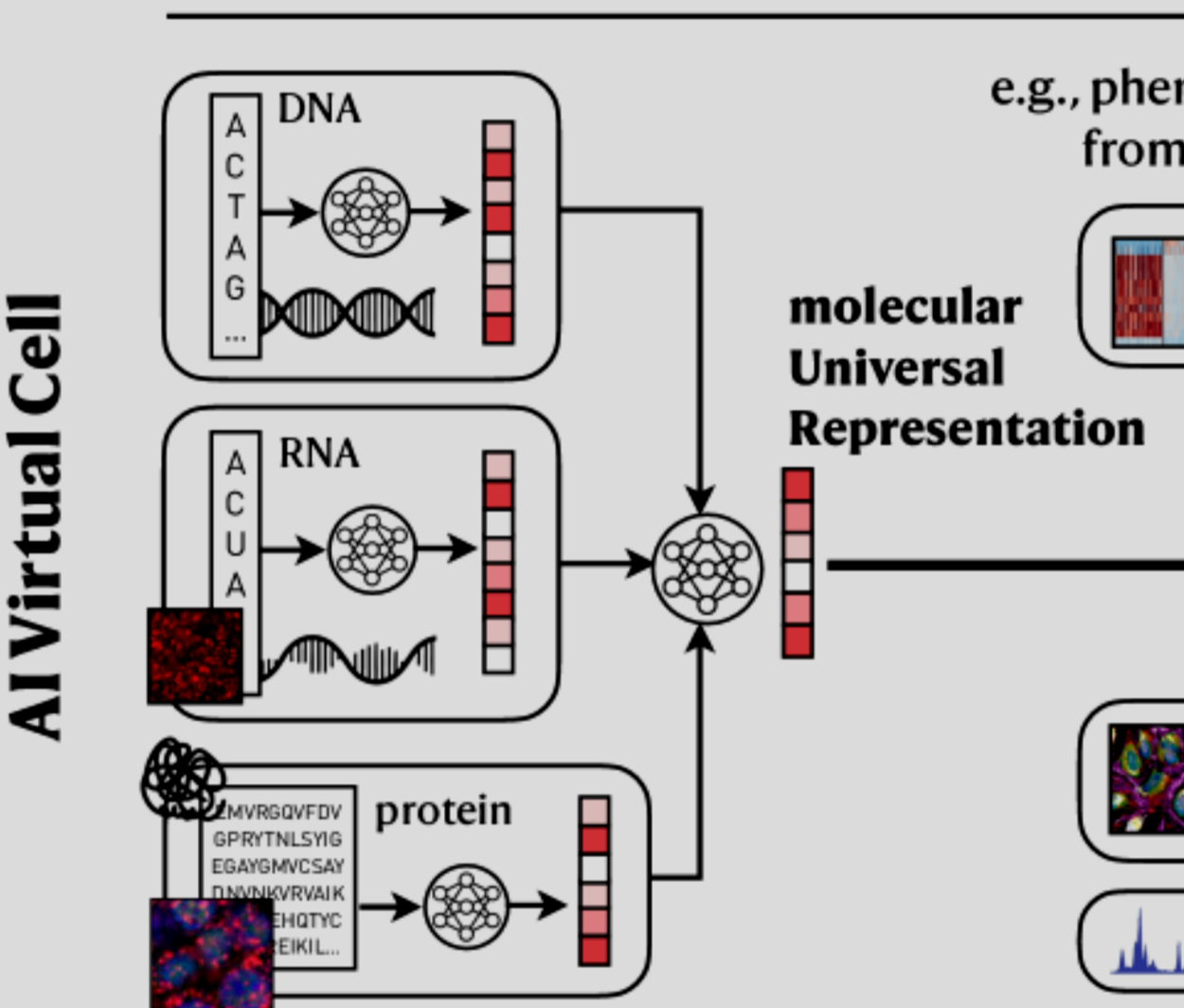
However, while working at Whitelab and even before that, I started to notice a change in the community. AI is slowly becoming more and more powerful and solving real biological problems. I saw it from foundation models in structural biology -with equivariant models and protein language models-, to transcriptomics and DNA language models.🤖🧬
During my work on celligner2 -at Broad-, I could already envision how foundational models would help in many aspects of my problem and many others. Working on the Atlas at Whitelab, they were always on our minds as possible solutions to tackle some real-world scientific problems (see my Whitelab post). (wait for a potential paper from our team at Whitelab in “nature computational science perspectives”) I had so many projects in mind and so little time to try to tackle them.💡📝
I also could see the speed of development of new papers and the breadth of possible research endeavors available to computational biologists. So there was some FOMO there for sure and I knew that if an AI-heavy transcriptomic research project was coming my way I might not say no.📈📊
So when I received this proposal from Laura and Gabriel I was very excited. It felt like the perfect fit for me. It had it all plus lots of applied math and stats plus gene regulatory networks which I spent years studying with Max Pimkin.💯📈
issues and realizations
But this is not the only reason: Before joining Whitelab and moving to France, I had interviewed at a few companies: BenevolentAI, DeepMind, Isomorphic Labs, Owkin, and DeepLife, and applied to many more. I could see for many of them that my not having a Ph.D. was an issue.
Other companies, like Whitelab or Aqemia, were very clear that my experience, to them was worth much more than what most Ph.D.s would bring.
But this really made me realize that life would be that much tougher without this diploma and that many doors would be closed or harder to open.🚪🔐
Finally, working in startup companies, I noticed their needs and challenges, some of them I believe specific to European startups:
The funding in Europe is often much smaller than in the US. This is problematic for biotech companies that need high CAPEX from the get-go to hire the best people, do experimental validation, build experimental facilities, and create a first pipeline. This is creating a state of slower initial growth than for their American counterpart.💰💼
This might seem to some like a small problem but it can become soul-crushing if you have been in the same company for years, waiting for it to enter exponential growth. It also can create issues, as the companies might lack important positions or teams with expertise that are harder to come by in Europe or need high investments.
Another realization I have had in both companies I was at, is the open-sourcing scare.😱📦
Open-source and creating value
Knowing biologists and Business Developers, I can guess this is an issue in many if not most (non-GAFAM) companies. “Use as many open source software and tutorials as you want”, “but don’t open source anything we do, don’t talk about anything we do”.
In software, since it is something that can get copied and pasted so easily, scarcity has to be artificial.
However open source and open science are often beneficial as everyone can collaborate and build on top of each other’s work. This is the inherent innovation and progress phenomena built into software development and research.🔍🌐
However, external pressures tend to force privacy towards innovation to keep it for oneself.🤐🔬
I would argue that for projects in which the result is very far off or unclear, not open-sourcing can be problematic. If you are building a product like a lamp, say. You cannot cheat, will the lamp turn on? For how long? For something like Facebook, even, you get usage metrics from very early on. For Biotechs, computational biology, and research, The clear-cut goal is often “a drug that passed some clinical trials”. This creates many untangible and not very engineering-like objectives along the way like: “Can you get people to believe in you?” how many partnerships do you enter?”.
In this context, you can lie. You can lie to the client but also to yourself. You can design a very nice-looking “car” that doesn’t drive at all. but since you are selling to people who don’t even really understand what is a car and what to do with it: it might be okay.🚗🤫 I am not saying that this is what we do at Whitelab at all! But it is an issue that all biotechs face and struggle with.I haven’t been the first to mention it.
But this is why I think open source is so important: when you let people know how to use a car, try your car, the iteration cycle is much faster. 📚🚗
In this context computational biologists should strive to inventivize environments where they can build things that will have a more direct impact than “helping make a drug”, through open source and through experimental validation.
This has been a big lesson learned for me too. Thinking more about impact, direct impact, and value creation. 🌍📈
the hard decision
Nothing was pushing me away from my current job and I felt welcomed, useful and that we were going forward in our grand vision See the Whitelab post. Choosing the Ph.D. really felt like choosing the hard thing, the hard path.
It is a hard decision because of how it might be seen too.
I so do not want people to see it as doing a U-turn, or as a start-over. I now have 5 years of experience in computational biology and I am afraid of hearing “you are just fresh out of your Ph.D. and we thus don’t think you are yet qualified for a lead role in our company.” from an HR manager after my Ph.D.📆👨💼
To me, it is a continuation of my career. Ph.D.s can be many things and I see mine as a kind of post-doc or a long sabbatical.
Moreover, staying for such a short period in companies is not seen well. For very sensible reasons. But startups are a special environment, especially so early-stage. I have stayed for 3.5 years at Broad. Much longer than most people in my position there. I have worked on my project, PiPle, for more than 5 years. I also think I have shown how much I cared for Whitelab and the personal dedication I have put in and continue to, in the company. 🌟🤝
Finally, I feel I have actually learned a lot more in these 10 months than I had in a long time. From structural biology and drug discovery to business and management, the startup ecosystem, and cell and gene therapies. This is really thanks to these fast-paced environments and all the people I have been lucky to work with.
the many lessons learned
With my previous experience in building a company, I would not have expected to learn that much from being in a startup. I think I did not realize the major steps that going from a company of 1 to 5 people is compared to 5 to 20 and 20 to 100, etc… each growth cycle is a full reinvention of the company culture, it is new challenges and things get exponentially more complex for the C-levels (i.e. the executive team: CEO, CTO, CSO, CFO, HR manager, etc)
being a great C-level in a startup
It shows how much of an example they are for everyone.
Here are the 4 points I learned for the C-levels:
- The C-levels’ domain knowledge is key to success. 💼🧠🔑
- The ability to create agile processes and to hire the right people at the right time is also key to success. 🔄👥⏰🔑
- Fostering a culture of trust and respect is how you can keep the best people in your company. 🤝💼
It might seem obvious but I really was shocked by the Impact of C levels on a startup culture. I was shocked also by the key “glue role” that C-levels need to play during this 1-100 people size. They are the connection between people inside the company and the partners outside it.🚀🔗
It shows how much of an example they are for everyone.
But the example they set needs is a constant balancing act between everyone’s needs and goals. Within all this chaos they need to keep fostering a culture of trust and respect. This “keep cool” in most situations is as hard as the balancing act they constantly play. 🤹♀️🤯
Finally, their domain knowledge 💼🧠🔑 is key to success. This requires experience in their topic and what their clients and the market want. All of this trickles down when they start hiring people: M.D.s, Ph.D.s, business developers, and engineers, with strong experience in their field are of the utmost importance in biotech. In here C levels again play a key role, who they are, the culture they have fostered allows them to recruit and keep the best people, especially when the company is still small and hasn’t reached a strong brand recognition yet.
Keeping good people is super tough: you need to pay them well, have their work be fulfilling while always showing them you are on a high growth trajectory. Processes and agility all play a role too. 🔄👥⏰🔑 Moreover, it is only through these well tought-out processes and structures that everyone’s strength and contributions can be channeled efficiently into the projects and products the company is building.
Thinking differently than in academia
Building a startup has many similarities with being a researcher, for example: you have ideas you need to sell through a powerful story. This is how you get people and funding. These ideas although incomplete have to convince people. You spend a lot of time finding money and building a team of experts and beginners.💡💰👥
But if it was that similar, every ex-PI would be a trememdous CEO. There are big differences too. The first few discussions with the BD team at Whitelab really made me understand them: First, you need to know at all times what you are selling and what people want, even though you might not have it yet. What counts is that you know you are able to have it and how you would do it.
Second, putting yourself out there: going to as many conferences and events, with a solution to people’s problems. This needs to happen from the get-go. (jeremie from the future here: I realize now that these 2 points are still things that many PIs would be familiar with haha)
Finally, your first product might be miserable, but if it solves someone’s problem, it’s okay. And thinking product first, even in comp bio, is not really a reflex. What is a product? when you see all these amazing databases, datasets, and tools. What does one bring? You don’t have to bring much initially, it just need to be useful to people. What is a product? It is really an abstraction over the work that your team does. It has to be polished, output meaningful results, within deadlines, it must have a given cost upfront… 🗂️📅📋
The future of cell and gene therapies
I joined Whitelab because I strongly believe in the future of genetic therapies. Covid opened our eyes to the power of mRNAs, while CAR-Ts have had a big impact on cancer therapies, and many more drugs are in the pipeline.
Especially understanding the issues that plague small molecules. It felt to me that we needed a more targeted approach for the medicine of the future.💉🧬🌐
But first, what are cell and gene therapies (CGT)s?
Cell and gene therapies (CGTs) represent a revolutionary approach to disease treatment, moving away from traditional methods towards more targeted and personalized solutions. These therapies work by altering the genetic material within a patient’s cells to fight or prevent disease. They can replace, remove or repair abnormal genes or introduce totally new genes to fight diseases.🧬💊🌡️
A very famous subcategory: mRNA therapies, such as those used in some COVID-19 vaccines, are a type of gene therapy that works by introducing a small piece of mRNA into the body. This mRNA carries some instructions and once inside our cells, it uses the body’s own machinery to execute it. The advantage of mRNA therapies is that they don’t alter the DNA in our cells and are not permanent, reducing the risk of long-term side effects.💉🦠🧬
Antisense oligonucleotides (ASOs) are another type of CGT that work by binding to the mRNA produced by certain genes and preventing them from being translated into proteins. This can be useful for diseases where a particular gene is overactive. ASOs can be designed to target almost any gene, making them a highly versatile tool in the fight against disease.💉🎯🧬
Overall, the main advantage of CGTs is their precision. They target the root cause of diseases at the genetic level, offering the potential for more effective and longer-lasting treatments with fewer side effects. This precision, combined with the rapid advancements in technology and our understanding of the human genome, is why many believe CGTs represent the future of medicine.💉🌍🔬
I believe that since genetic sequencing and machine learning, we are able to understand the root cause of disease in the name of one or a few cell types and their disregulation caused by often one or a few genes.
Being able to target only these specific diseased cell types is within reach for CGTs but not for small molecules. Moreover being specific in what we are doing: what genes we are triggering is also what CGTs can do that most small molecules can’t.
It feels like a perfect CGT would allow us to create a novel drug in hours. From a laptop and a DNA printing machine. It would be the perfect interface between man and life.
My goals and objectives for the Ph.D.
A main mistake during one’s Ph.D. is to not see the time passing by. My goal for this Ph.D. is to be as product-first as I was at Whitelab. Delivering results quickly & improving until it is publishable.🏆📊📚
This mistake, thinking “Well I have 3 years…” is what makes people go overboard, finish stressed, and unprepared for what is next. Thus I plan for doing mine in 2 years. And I will prepare everything around this idea. I will also start to think about what is next ASAP.
To do that best, one needs to take the chance of the Ph.D. to make connections with other labs (industry or academic) on my end I am thinking for example at Broad, the Theis lab, the Yosef lab, Valence labs, …🤝🔗
Moreover, a good advice I have been given is to know what you want to do and what you don’t want to do. Know what you are here for. Learn to say no. And I learned to say no in the last 4 years. My goal is to work on large models & large datasets, mostly transcriptomics. But also making sure to always go back to first principles and biology.🧬🧪🔍
I also know I want to make something useful, make something that can be a stepping stone for others. Something that has an impact on the community. I know that to do that you have to go the extra mile in terms of dev with and be honest with yourself about any shortcomings.
Finally, I have been very lucky to often become addicted to my work. I like working hard and I like challenges. But for this to happen, I need to keep enjoying what I am doing. I also wish to have no regrets about this decision.
Thus my final goal is to enjoy it, as much as I can!😊🚀
Recap
Thus my motto for this Ph.D. will be:
- do it in 2 years and be prepared
- make as many connections as I can
- maximize impact on the community: make something useful
- enjoy it as much as possible
Ph.D. proposal
Link to the proposal
Previous proposals
I have had a previous Ph.D. projects proposals for a Ph.D. that I then postponed. I list it here too for historical purposes.
]]>